Current and Previous Research Projects
|
Visit us at our new webpage HERE for details about ongoing research in the Skeletal Genomics lab at the Max Planck Institute for Evolutionary Anthropology! |
|
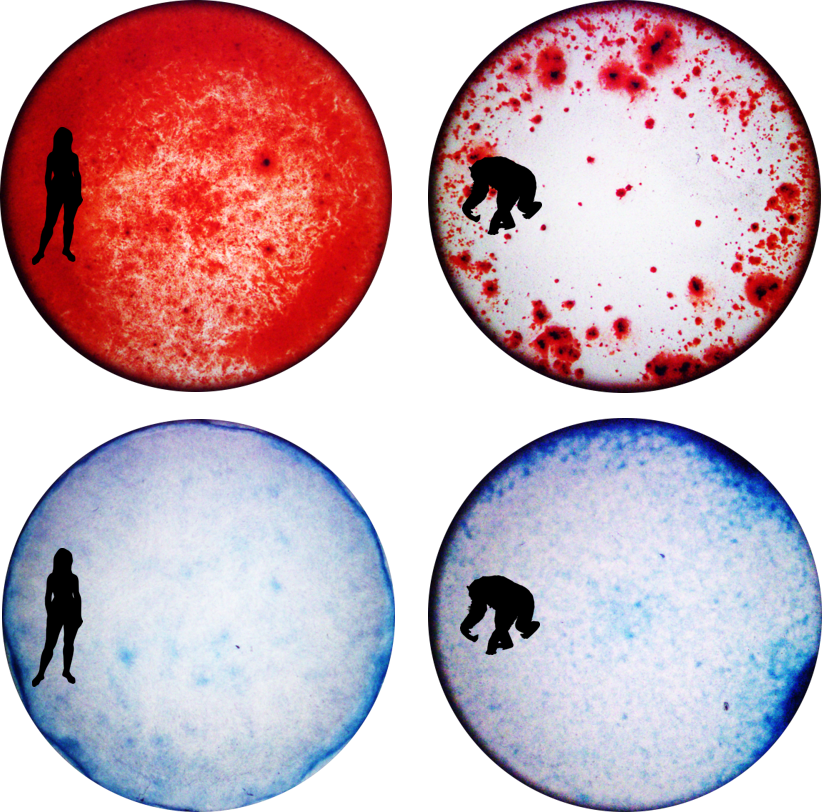
iPSC-derived osteoblasts (top) with Alzarin Red staining of calcium deposits and iPSC-derived chondrocytes (bottom) with Alcian Blue staining of collagen extracellular matrix in a human cell line (left) and chimpanzee cell line (right).
(Image Source: Genevieve Housman) |
Comparative Primate Skeletal Cell Culture ModelPrimates display diverse skeletal morphologies and susceptibilities to skeletal diseases. However, due to the limited availability of skeletal samples appropriate for molecular research, there are relatively few studies that characterize molecular mechanisms associated with complex skeletal phenotypes and even fewer studies that consider the effects of gene by environment interactions in a population or evolutionary context.
In order to examine gene regulation variation in skeletal cell types, assess how these patterns differ within and among primate species, and determine how environmental forces like biomechanical strain impact cellular phenotypes, their susceptibilities to disease, and their capacities to recover from disease, I have developed a comparative primate skeletal cell culture model. This system consists of several human and chimpanzee induced pluripotent stem cells (iPSCs) that are first differentiated into mesenchymal stem cells (MSCs) and then subsequently into osteogenic (bone cells) and chondrogenic (cartilage cells). I am additionally exposing skeletal cell types to mechanical strain and inflammatory cytokine treatments that serve as an in vitro model of osteoarthritis (OA) in order to determine what species-specific regulatory responses may make chimpanzees less susceptible to or more protected against OA than humans. This project was part of my postdoctoral research at the University of Chicago. |
|
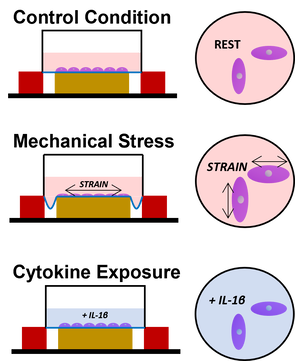
in vitro OA treatments used to examine dynamic gene regulation responses at population- and evolutionary scales.
(Image Source: Genevieve Housman) |
Gene-by-Environment Interactions in the SkeletonSkeletal cells are highly responsive to environmental signals. At a cellular level, responses may include cell differentiation, extracellular matrix production, or hypertrophy. At an anatomical level, responses may include growth, bone remodeling, or pathogenesis. Responses to environmental stimuli may be mediated through gene regulation changes which may be partially genetically controlled. However, such gene-by-environment interactions are not well characterized in primate skeletal cells.
Using my skeletal cell culture model, I am more deeply interrogated the mechanisms by which genetic variants and environmental effects interact to alter gene regulation and impact complex skeletal traits. I am currently examining perturbations that result in cells developing a matrix degrading phenotype similar to that observed in osteoarthritic (OA) tissues - mechanical stress treatments and inflammatory cytokine exposures. In a large, population-scale panel of human iPSC-derived skeletal cells, I aim to identify mechanical stress response expression quantitative trait loci (eQTLs) that are relevant for understanding OA pathogenesis, and in a comparative, evolutionary-scale panel of human and chimpanzee iPSC-derived skeletal cells, I aim to identify interspecific transcriptional responses that may provide additional insight into OA susceptibility differences across species. This project was part of my postdoctoral research at the University of Chicago. |
|
Extracting DNA from nonhuman primate bone and cartilage samples. (Image Source: Genevieve Housman)
|
Primate Skeletal DNA Methylation VariationPrimate species share many genetic features but differ in their appearance and behaviors. Epigenetics can help explain this discrepancy. The epigenome comprises mechanisms that act above the genome to regulate gene expression. DNA methylation is one such mechanism in which the binding of methyl groups to DNA silences gene transcription and the removal enables transcription. Variation in DNA methylation patterns which produces differential gene expression can contribute to phenotypic variation. Nevertheless, studies of nonhuman primate DNA methylation have been limited to a small number of tissue types in a few species. In particular, hard skeletal tissues have not been readily examined.
The aim of this study is to identify DNA methylation in nonhuman primate skeletal tissues and assess variation both within and between species. This research is the first of its kind, and results will further develop our understanding of primate epigenetic variation. These data will also help to build a phylogenetic foundation on which ancient hominin skeletal epigenetics can be compared. This project was part of my doctoral dissertation research and was conducted in collaboration with Dr. Ellen E. Quillen at Texas Biomedical Research Institute. |
|
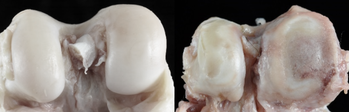
Healthy baboon knee cartilage (left) and severe osteoarthritic baboon knee cartilage (right). (Image Source: Lorena M. Havill)
|
Evolutionary Epigenetics of Osteoarthritis
According to the WHO, the degenerative joint disease of osteoarthritis (OA) is present in 9.6% of men and 18.0% of women ages 60 or older world-wide and results in severe movement limitations. The CDC further notes that knee OA is especially prevalent in the U.S. and results in over $28.5 billion of hospital expenditures. Thus, it is important to understand the causes of OA.
Epigenetic factors are now thought to play a more influential role than genetic and environmental factors alone, but few epigenetic studies have taken an evolutionary perspective. To address this issue, this project assesses the relationship between skeletal DNA methylation patterns and knee OA development in baboons. These animals serve as a primate model of OA. Findings in baboons are compared with previously published human data to better understand the phylogenetic context of OA pathogenesis. This work has wide implications for the field of evolutionary medicine as it attempts to evaluate how epigenetic-disease relationships have evolved in the primate lineage. This project was part of my doctoral dissertation research and was conducted in collaboration with Dr. Ellen E. Quillen at Texas Biomedical Research Institute. |
|
Measuring 3D landmarks on baboon femur bone using a Microscribe (Image Source: Genevieve Housman)
|
Epigenetics and Skeletal Morphology
Primates have diverse skeletal anatomies that evolved to fit many forms of locomotion, and such variation is often used to reconstruct extinct primate species and to support theories on primate phylogenetic and evolutionary history. However, the mechanisms contributing to these hard tissue phenotypes are still unclear. Genetic and environmental factors have some impact, but the role of epigenetic factors has not been fully evaluated in nonhuman primates.
This research aids in this endeavor by identifying how DNA methylation patterns in primate skeletal tissues relate to femur morphology both within and among several nonhuman primate species. This work attempts to bridge the gap between molecular mechanisms and morphology in order to answer evolutionary questions about how epigenetic factors contribute to complex skeletal phenotypes in primates. This project was part of my doctoral dissertation research and was conducted in collaboration with Dr. Ellen E. Quillen at Texas Biomedical Research Institute. |
|
Besides humans, armadillos are the only other known host of leprosy. (Image Source: Richard Truman)
|
Evolution of Mycobacterium in Animal ReservoirsZoonotic mycobacterial infections continue to impact modern human populations. Thus, methods able to survey the presence of mycobacteria in potential animal hosts is necessary for proper evaluation of human exposure threats. This project tested such methods in a trial study of armadillos artificially infected with Mycobacterium leprae, the pathogen that causes leprosy. Detection of mycobacterial-specific single- and multi-copy loci using quantitative PCR proved to be a viable method for such surveys. Thus, this technique was subsequently used to detect mycobacteria in wild marmosets.
This research was initiated during my masters work at Arizona State University. Project collaborators include Dr. Richard Truman, Dr. Joanna Malukiewicz, and several researchers in Brazil. |
|
Cancer cell lines before drug treatment. (Image Source: Genevieve Housman)
|
Cancer Epigenetics
Cancer development, classification, and treatment are heavily researched areas, but many debates in this field remain unresolved. The association of genetic and epigenetic markers with particular cancer types has advanced the field of personalized medicine. However, there may be global epigenetic changes that contribute to carcinogenesis, metastasis, and the development of drug resistance. Targeting these pathways in combination with specialized treatments may be more effective than either option alone. This research began to test this idea by assessing the effects of epigenetic drug treatments and other drug treatments, separately and in combination, on cancer cell lines.
This work was initiated during my undergraduate studies at Boston University. Project collaborators include Dr. Sibaji Sarkar and several former students from this laboratory. |